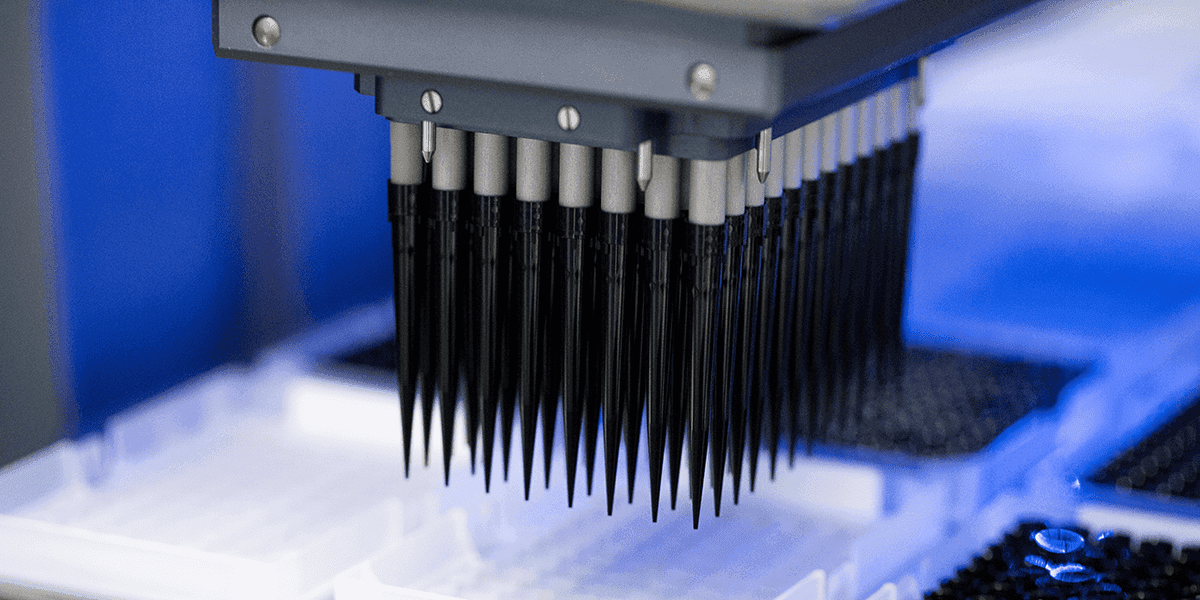
The Exome+® Assay
Get the performance of a targeted panel, the breadth of a microarray, and the completeness of an exome — all from one assay.
Avoid compromise
Through custom development work and proprietary bioinformatics solutions, we’ve created an assay that meets the needs of both clinicians and researchers in contrast to off-the-shelf assays that are typically optimized for one or the other.
Read on to learn how we’ve eliminated the compromises clinicians and researchers typically have to make on cost, quality, and breadth when choosing an assay.
An Exome, plus more
A panel-grade, clinical exome plus a whole genome backbone enabling tens of millions of imputed variants as well as the full mitochondrial genome.
Exome+ powers our Traits analysis, which includes fitness, diet, health, and wellness content. This comprehensive approach allows us to provide both clinical-grade results and personalized insights.
Panel-grade
A ≥ 99.5% call rate across ~ 600 genes that are important in the areas of hereditary cancer, cardiovascular disease, pharmacogenomics, and carrier screening.
Comprehensive
Covers ~20,000 genes and detects and reports SNPs, indels, multinucleotide variants, substitutions, and complex variants, intragenic copy number variants (CNVs), and multigenic CNVs.
Key performance metrics


Why call rate matters
Average coverage isn’t sufficient
Many labs often describe an assay’s coverage primarily in terms of the mean coverage across the entire assay, but that only tells part of the story.
An assay’s coverage can vary greatly within a genome, exome, or panel. For example, regions may have reduced coverage due to high or low GC content or because other similar sequences reduce the mapping quality of the reads.
As a result, a test with an average of 100x coverage may have large regions with no coverage at all even though the assay’s overall average is high.
Call rate helps to identify gaps in coverage
What is important to understand about the performance of an assay is the percentage of the region that receives trustable results (known as the call rate). Looking at base-by-base call rates can reveal coverage gaps, even in assays with high average coverage.
We optimize the Exome+® assay for uniform read distribution across regions that are important for specific types of genetic testing. For these genes, we provide high-confidence calls at a rate of ≥99.5%, with base-by-base coverage at >20x coverage.
Combining the advantages of several assays into a single one.
Helix’s Exome+® assay has been carefully optimized to provide the benefits of lcWGS, whole exome sequencing, microarrays, and targeted panels — all in one assay.
Targeted Panels
Panels are often used in clinical settings because they're low cost and provide high quality coverage of specific, pre-defined genomic regions of interest. While useful in certain settings, such assays are limited by their narrow focus.
With ≥ 99.5% call rate across ~600 genes relevant to many disease states, the Exome+ assay enables you to run multiple panel-grade analyses with a single sample.

Whole Exome Sequencing
Standard whole exome assays offer broader coverage (compared to targeted panels) but are known to be incomplete and to suffer from gaps in coverage. Because of this, exome sequencing is generally inappropriate for clinical applications that require panel-grade testing.
Helix’s Exome+ assay maintains a consistently high call rate thanks to a uniformity of coverage across all coding regions.

lcWGS and microarrays
The Exome+ assay harnesses the utility of lcWGS and microarrays by covering a whole genome backbone enabling genome-wide imputation and ancestry analysis. Imputation accuracy of the Exome+ assay is comparable to 0.5x lcWGS.

A future proofed assay
There are several advantages to running panels in the presence of a whole exome, including flexible test design, flexible re-testing, and increased functionality.

The Exome+ assay provides the flexibility needed to build a comprehensive panel that can be expanded over time. With ~20,000 genes sequenced in each sample, every gene becomes a candidate for inclusion. And, while we have ensured the highest call rates across those genes that are most likely to belong to a panel, the Exome+ assay delivers high call rates for all genes beyond this core set.
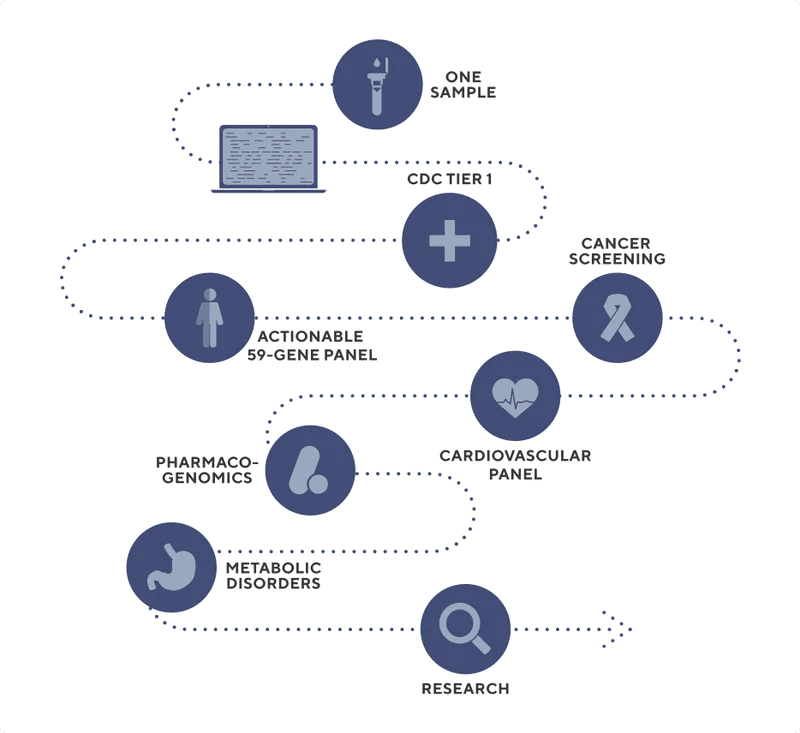
The Exome+ assay supports many use cases off of a single assay. This means that once an individual has been sequenced and tested, the data is available and can be retested for additional use cases as needed. This enables health systems to lower costs while improving workflows and patient engagement.

Typical commercial pharmacogenomic assays only report on the most common star alleles, leading to inaccurate results for genes like CYP2D6 in over 15% of samples. The Exome+ assay avoids this through proprietary bioinformatics solutions that enable reporting on all known CYP2D6 star alleles (106) with 100% accuracy, making it the most comprehensive and accurate CYP2D6 calling on the market. Helix’s pharmacogenomic solution additionally reports on all known star alleles for CYP2C19, CYP2C9, and more.

Ask specific questions, get specific answers
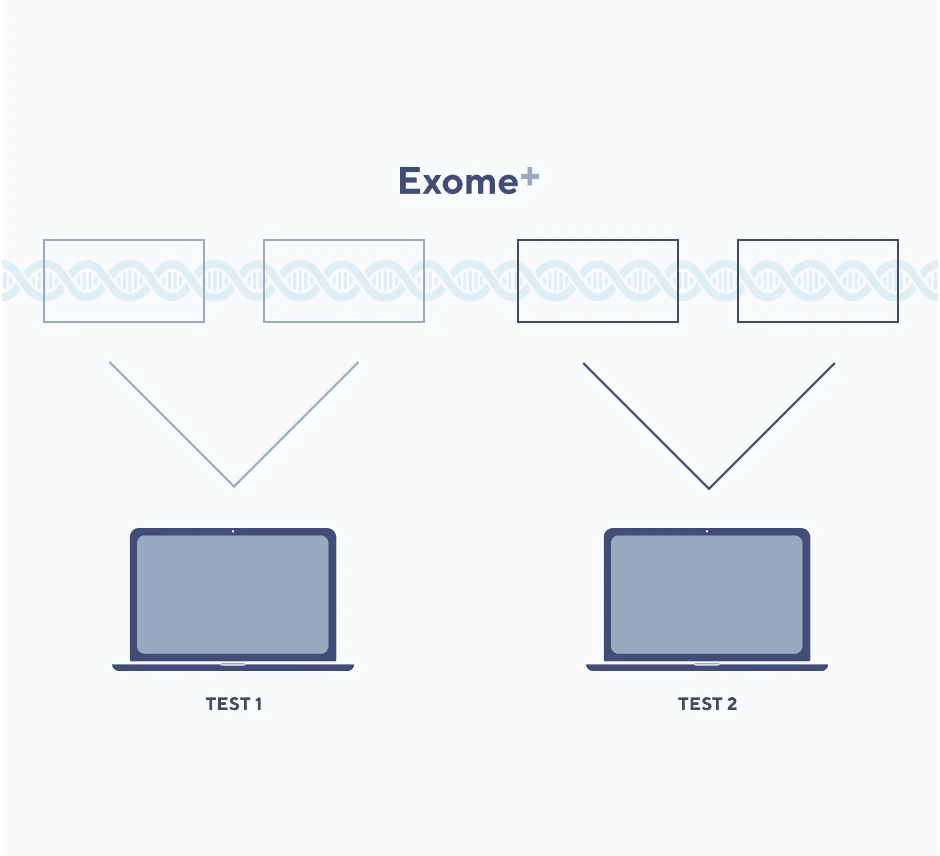
Focused data analysis
Helix securely stores the entire Exome+ data set and only “unlocks” the genomic data you request. Data is made available for interpretation via targeted bioinformatics queries. All other data remains uninterpreted, mitigating concerns about incidental or secondary findings (knowledge about potentially impactful findings outside of the interrogated data).

