If you’ve ever taken a biology class, you probably learned about Punnett squares—the tic-tac-toe-ish diagrams that help you estimate the likelihood of a specific trait being passed onto the next generation. Commonly, this is illustrated using eye color. The idea here is that brown eyes are dominant over blue eyes, and that you have to get two copies of the blue-eyed variant to have blue eyes yourself. While this concept has been taught for decades, it’s not entirely true.
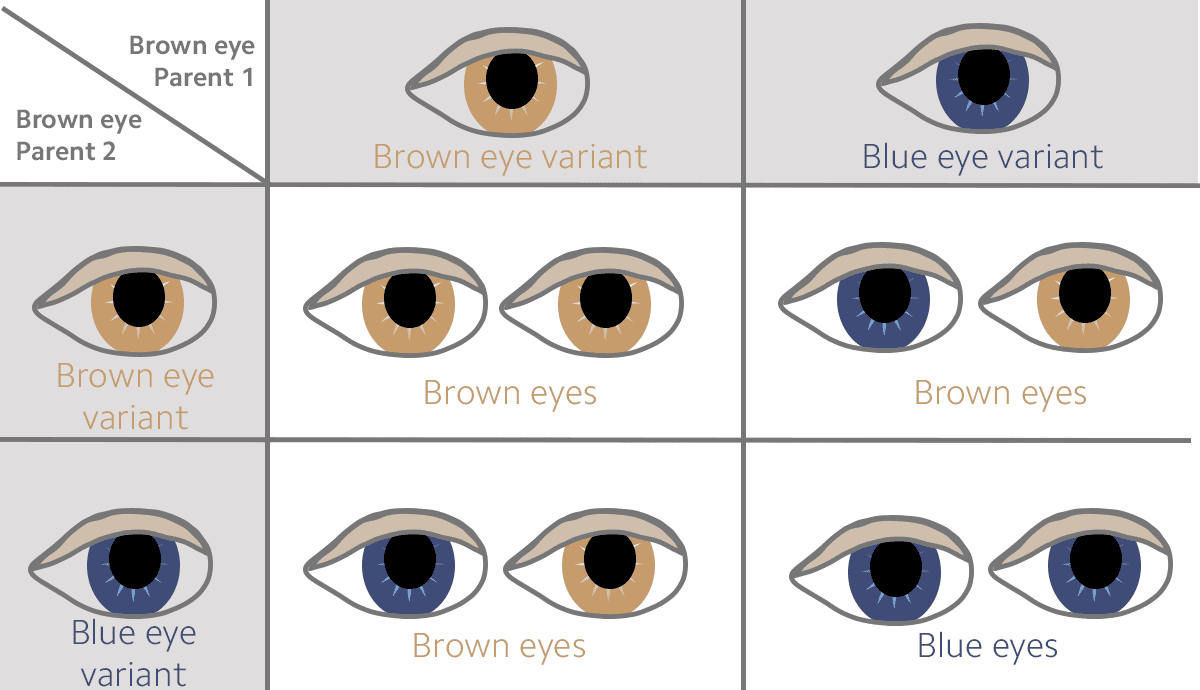
This Punnett square for eye color shows that the brown eye variant is dominant to the blue eye variant, just like you may have learned in school. In reality, eye color is more complicated than just two variants.
The idea of one gene being responsible for one trait (like eye color) is overly simplistic. In reality, we know that over a dozen different variants in your DNA contribute to eye color. It’s true that some variants are more important than others, but you won’t get the most accurate estimate of eye color unless you consider them all. For example, variants in the HERC2 gene play a large role in determining whether your eyes are light or dark and is mostly responsible for blue eye color. But you won’t know if someone’s eyes are blue or grey or even green without looking at additional variants. Considering more than one variant at a time has a name: it’s called a polygenic score (PGS).
What is a polygenic score?
A polygenic score is simply an algorithm—one that adds up the impact of multiple variants. A PGS can analyze just a few variants, or it can consider millions. The key is that these variants have to be considered in aggregate. Some traits are based on just the presence or absence of a single variant, like whether you can taste a specific bitter compound called PROP or whether you have cystic fibrosis. But the vast majority of traits, from height to cardiovascular disease and vitamin D levels to male pattern baldness, are influenced by many variants and are therefore best estimated with this type of approach.
Let’s take the example of height. Height is determined almost entirely by your genetics. However, variants with the largest effect only add about 2 centimeters to someone’s height. If I told you someone had just one of those variants, you wouldn’t really know how tall they are. But what if I told you that someone had 1,000 “tall” variants while most people in the population only have about 800? You’d probably start having a better idea that they may be taller than average.
And that’s an important point about polygenic scores—they help put one person’s result in the context of the rest of the population. Knowing how many tall variants you have is only interesting if you can see where you fall in the population.
Why is it important?
So why should you care about polygenic scores? Well, you may already be familiar with them—after all, ancestry calculations based on genetics are a type of polygenic score, incorporating hundreds of thousands of individual variants. On top of that, PGSs are becoming increasingly common, thanks in no small part to larger studies and the trend to collect phenotype data (e.g., UK Biobank). This makes identifying variants associated with certain traits much more likely. In fact, Myriad, a molecular diagnostic company that offers genetic tests for inherited breast cancer, recently announced that it has incorporated a PGS into its reports on breast cancer risk.
Which brings me to the second reason PGSs are important: they’re changing how we think about risk. Previously, if a woman didn’t have a variant in known breast cancer genes, there wasn’t much we could tell her about her breast cancer risk, other than by looking at her family history. However, if a polygenic score is available, every woman can receive risk information based on this PGS. This approach may allow everyone to have a better sense of their own genetic risk for certain traits or diseases than just using family history alone. In fact, PGSs are sometimes called polygenic risk scores due to the approach becoming popular in disease contexts.
What’s next?
If PGSs are already being used, where do we go from here? Almost certainly we are going to see improvements on how scientists and researchers calculate these scores. New statistical methods will be tried and validated. And as our datasets get larger and are linked with richer data from electronic medical records and diverse biobanks like GTEX and the UK Biobank, we are likely to continue discovering additional traits that can benefit from a PGS approach. Eventually, we will likely see scores that use many genetic variants, as well as different environmental variables as well.
As next generation sequencing (like Helix uses) continues to grow, we will likely see these scores become more predictive, due to the discovery of rare variants with larger effects. Previous efforts have focused on common variants (those present in at least 5% of people) because they rely on genotyping. But sequencing will allow scientists to dig deeper into the genome, identify more relevant variants, and improve the power of the PGSs.
We already have many tools for deriving an incredible spectrum of important insights from the human genome, but polygenic scores hold the promise of broadening that spectrum. Whether it’s helping us understand the genetics behind human height or giving us the ability to calculate a person’s risk of developing complex diseases, PGSs are likely to play a large part in the future of genomics.
