Early humans relied on word-of-mouth stories—passed on from generation to generation—to learn what the world looked like in years past. Once invented, writing became the “time machine” of choice; Aristotle’s words help us see what people thought more than 2,000 years ago, for example. But what if we want to know what life was like before written text, before the beginning of modern oral history? How can we see the world that existed more than 50,000 years ago? For this, we turn to the stories contained within our DNA.
For millions of years, DNA sequences have been evolving in the same way that an urban legend might evolve with each new telling. Imagine for a moment that you overhear two people swapping stories: One talks about the North American Bigfoot as a 6-9 foot tall ape, the other talks about the 1.5m tall Canadian Nuk-luk bush-man. The stories are nearly identical except for the height of the mysterious creature and some wording. Later, when you recount the story, you might mix up the height difference and just report that the Canadian Nuk-luk is a 9 foot tall ape. This hypothetical example demonstrates two real concepts in genetics that help us see into the past.
The first concept demonstrated here is known as recombination. All humans have a nearly identical DNA sequence that will, on average, match at about 99.9% of locations. This is similar to how the story of the Bigfoot is almost identical to the Nuk-luk with just a few details differing. Inside each of us, there are two sets of DNA—one from each biological parent. Each set is nearly identical except for subtle differences. Before your DNA can be passed onto the next generation, it will go through the process of recombination. This is where the two sets of DNA line up next to one another and randomly swap similar segments of DNA. In terms of the urban legend, recombination is like the Bigfoot’s height being swapped with the Nuk-luk’s. The end result is a hybrid DNA sequence that contains segments of DNA from each of your biological parents. This DNA swap can happen in most parts of the genome (with some exceptions), and contributes to maintaining genetic diversity within the human population.
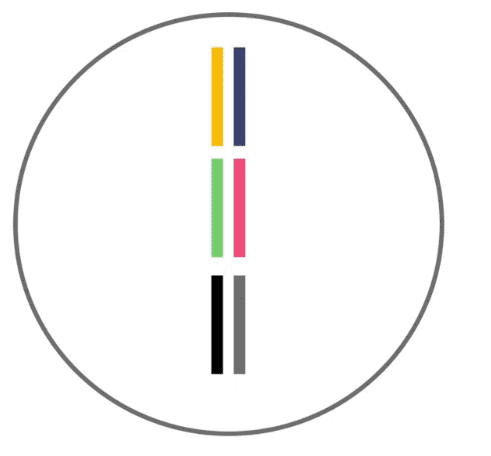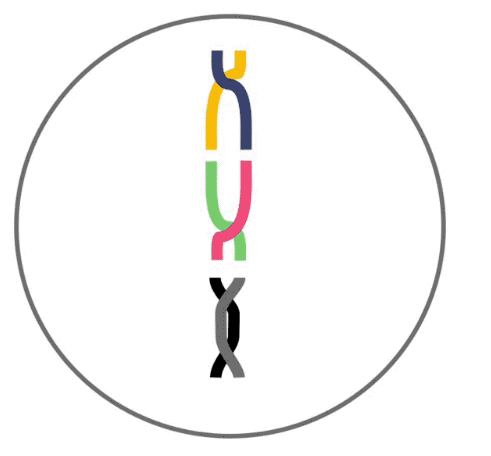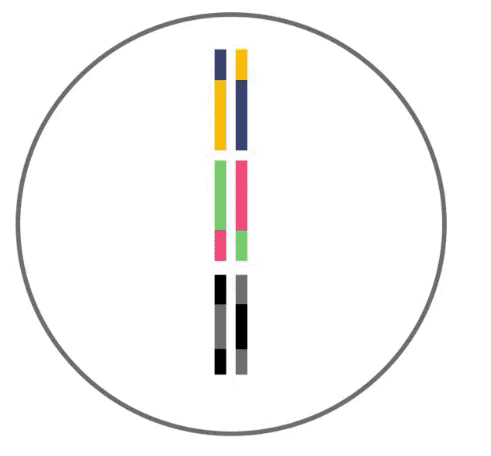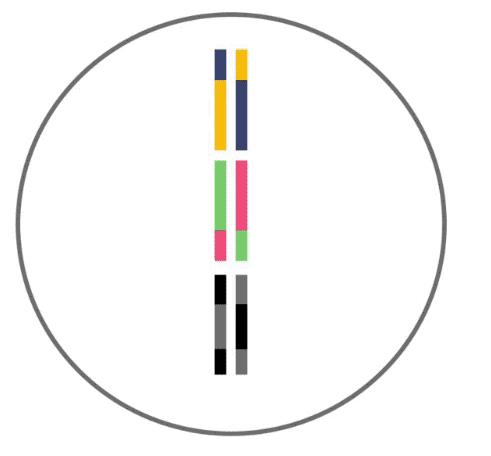
Here, recombination is demonstrated. Recombination like this occurs before your DNA can be passed to the next generation. Three segments of DNA from one parent (shown as yellow, green and black lines) align next to three segments from the other parent. DNA is then swapped between segments in random locations to produce hybrid DNA sequences.
The second concept demonstrated is linkage disequilibrium (LD). Though the name is a bit intimidating, the concept is fairly simple: LD is where two or more segments of DNA tend to be inherited together because they are commonly found together within the population1. In terms of the urban legend, notice how the numbers describing the creatures’ heights weren’t the only thing swapped—the units describing the height were also swapped. If information between the stories is being traded randomly, you might expect to hear an equal number of subsequent stories where the Nuk-luk is described as being 9 meters tall, 9 foot tall, or 1.5 feet tall. But if in a population of people, you mostly see it described as being 9 feet tall, then you would conclude that the number “9” and the units “feet” are dependent on each other in some way. We could also say that the number and units are in linkage disequilibrium. Scientists would then wonder why they have traveled together—it could be that no one would believe a story about a 9 meter tall animal, or it could simply be that the story hasn’t been told enough times for someone to mix up the units. Whatever the reason, it’s clear that—for the time being—the numbers and units are in linkage disequilibrium within this population.
Why does all of this matter? It can help us look back in time
Why does all of this matter? Understanding how often recombination takes place, and the statistical likelihood of two genes being seperated, can help us look back in time.
About 100,000 years ago, there was more than one type of human walking around on the planet2. Short and stocky humans known as Neanderthals and Denisovans were occupying the Eurasian continent. In Africa, it was the taller, lankier Homo sapiens. At some point, these two human groups met. What happened next has been the topic of much debate. Archeological evidence tells us that Neanderthals disappeared around 40,000 years ago2. But what caused this? Did Homo sapiens wipe out the Neanderthals through warfare, or did both groups simply merge? To help answer this, researchers have turned to genetics and studying linkage disequilibrium.
DNA sequencing of people around the world has shown us that about 2-6% of the DNA in people of non-African heritage comes from Neanderthals or Denisovans. We know this because researchers have compared the DNA sequence of ancient Neanderthal and Denisovan remains with those of Homo sapiens. In this data, they found patterns where DNA sequences were commonly found together, and that these patterns made a genetic signature unique to each population. Portions of these sequences from Neanderthals and Denisovans can be found in the genomes of many modern-day humans. Knowing the statistical probability of these sequences breaking up by recombination when they coexist with other sequences, scientist can infer that these sequences were initially in other populations and they can calculate how much time the Neanderthal’s sequences have spent in Homo sapiens populations3.
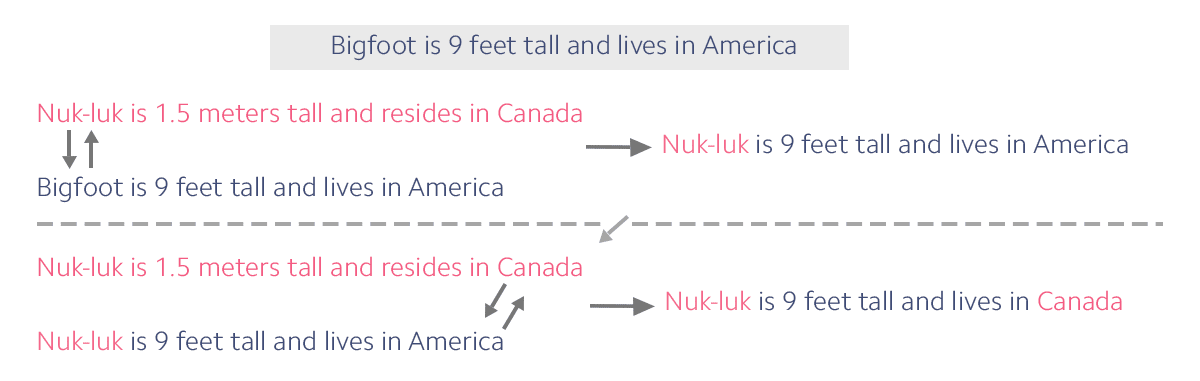
Consider the example shown here in which someone brings the story of the Bigfoot into a new culture where people only know the Nuk-luk story. Watch as it changes over time.
In this first panel, you can see that in just one generation, only a limited amount of information can be swapped. First, it was the creature’s name. Then, the words “Canada” and “America” were swapped. With this predictable limit on the amount of information that can be swapped, we can determine how likely it is that any one word will be separated from another by recombination. Over time, it becomes less likely that the block of story from the original Bigfoot legend will stay intact.
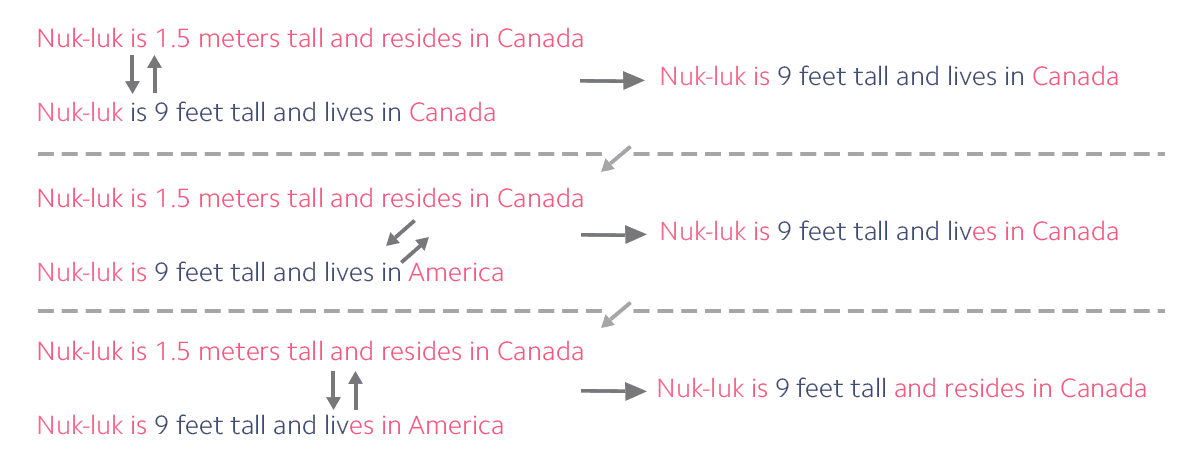
After multiple generations, the story has changed significantly, but still retains a block from the original Bigfoot story. Now, if a researcher were to find any of these stories, they could compare it back to the originals. If a large amount of the original Bigfoot block is still intact, they can be reasonably sure that the two different stories came together only one generation ago. But if the Bigfoot block is very small, they can do the math to figure out how long ago the stories met. This is how researchers calculated when Neanderthal and Denisovan DNA merged with Homo sapiens DNA. (In reality, the process is much more complex, but the theory is the same.)
Researchers have determined that Neanderthal and Denisovan DNA entered the genome of Homo sapiens sometime between 40,000 and 65,000 years ago. This provides evidence to suggest that some Neanderthals may have been absorbed into Homo sapiens culture and that warfare is likely not the whole story behind their disappearance. This kind of analysis isn’t just limited to Neanderthals—scientists use this method to learn about many events in human history, including the complex dynamics of ancient human groups in Africa, or the more recent interactions between migrating human groups across the globe2,3.
1Slatkin, Montgomery. “Linkage Disequilibrium — Understanding the Evolutionary Past and Mapping the Medical Future.” Nature reviews. Genetics 9.6 (2008): 477–485. PMC. Web. 5 July 2018.
2Nielsen, Rasmus et al. “Tracing the Peopling of the World through Genomics.” Nature 541.7637 (2017): 302–310. PMC. Web. 14 July 2018.
3Sankararaman, Sriram et al. “The Date of Interbreeding between Neandertals and Modern Humans.” Ed. Joshua M. Akey. PLoS Genetics 8.10 (2012): e1002947. PMC. Web. 14 July 2018.
